Genetics Based Computing
A Biologically Modeled Computational Paradigm
Nature often solves information problems in ways that are both elegant and efficient. Biological systems have evolved methods for storing, transmitting, and transforming information that operate reliably across billions of organisms and millions of years.
One of the most remarkable examples of this is genetic encoding.
Living organisms store enormous amounts of structured information inside molecules of DNA. This information determines the structure of proteins, governs cellular processes, and ultimately defines the physical form and behavior of an organism.
This raises an interesting question.
If nature has already developed a powerful information encoding system, could similar ideas be applied to computer science?
In other words, could a computational architecture modeled on genetic encoding offer advantages over traditional binary systems?
This article explores a conceptual framework for such a possibility.
Brief Summary of DNA Structure
DNA stores information using four nucleic acids, often called bases:
Adenine (A)
Thymine (T)
Cytosine (C)
Guanine (G)
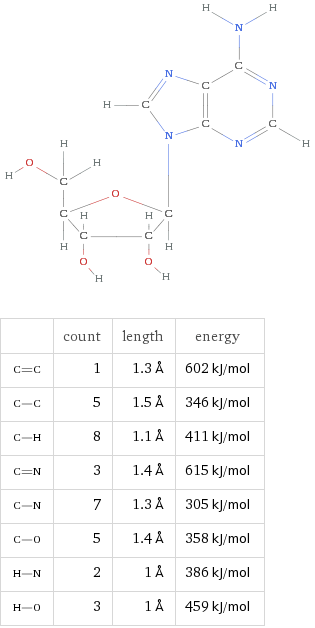
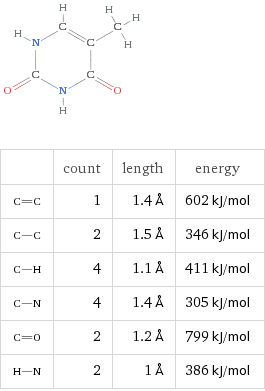
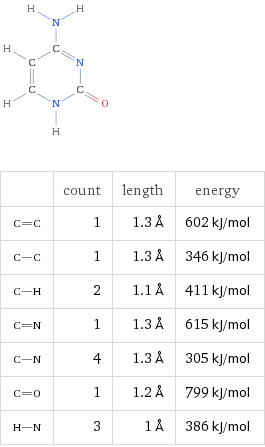
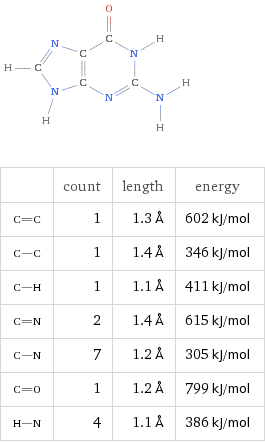
These bases pair according to simple geometric rules:
- Adenine pairs with Thymine
- Cytosine pairs with Guanine
These combinations are known as base pairs.
The sequence of these pairs forms the genetic code used by biological systems to store and process information.
Binary Computing vs DNA Encoding
Modern computer systems rely on binary logic, where each bit represents one of two possible states:
0 or 1
Groups of eight bits form a byte, allowing 256 possible values.
DNA uses a different approach.
Instead of two states, DNA uses four possible symbols:
A
T
C
G
From a computational perspective this is equivalent to a quaternary number system with four possible values per position.
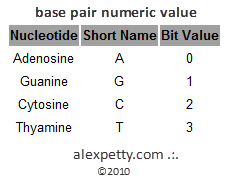
This difference leads to an interesting comparison.
A standard binary byte uses 8 bits to encode 256 values.
DNA uses three base positions, known as a codon, which produces:
4³ = 64 possible combinations.
While the raw capacity is smaller per codon, the structure of DNA encoding introduces other advantages such as redundancy, error correction, and self-organizing pairing behavior.
Binary Byte Example
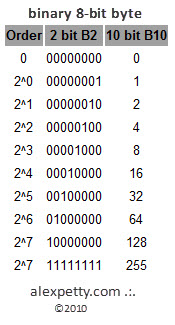
Binary systems represent numbers as powers of two.
For example:
73 in binary:
01001001
which corresponds to
64 + 8 + 1
Quaternary Codon Example
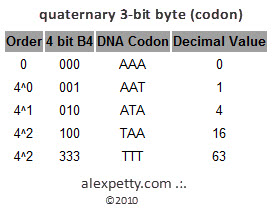
A quaternary representation uses powers of four.
For example:
53 in base 4 is
310
which corresponds to
3 × 4² + 1 × 4¹ + 0 × 4⁰
In a DNA-like encoding this could be represented by a codon sequence such as TGA, which would pair with ACT.
Toward a DNA-Inspired Encoding System
If digital information were encoded using a quaternary system similar to DNA, it might be possible to design storage and compression methods that take advantage of complementary pairing and structural redundancy.
One idea would be to store only one side of each complementary pair, allowing the matching half to be reconstructed algorithmically.
In principle this could reduce storage requirements and enable new forms of compression modeled on the double-helix structure of DNA.
Conceptual Compression Model
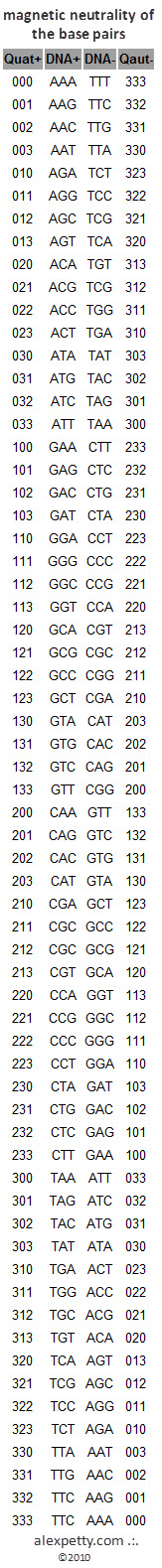
In this conceptual model, base pairs are treated as complementary numeric structures.
Only one half of each pair needs to be stored explicitly, while the other half can be inferred.
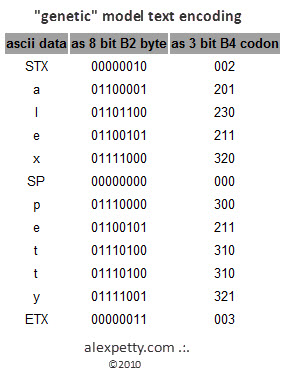
This produces a theoretical efficiency improvement compared to simple binary storage.
The compression ratio would depend heavily on the encoding scheme and the redundancy of the input data.
Runtime Translation Layer
Because modern hardware operates in binary logic, a DNA-inspired encoding system would require a translation layer.
Such a runtime system would convert between:
Binary hardware representation
Quaternary DNA-style encoding
This translation could be implemented at the software level until hardware capable of native multi-state logic becomes practical.
Optical Input and Output
An interesting parallel exists between quaternary encoding and color systems used in printing and imaging.
The CMYK color model uses four states:
Cyan
Magenta
Yellow
Black
In principle, optical systems could encode or transmit quaternary information using combinations of these colors.
Interestingly, chromosomes themselves exhibit color variations under certain forms of microscopy.
While this observation does not imply a direct computational role, it highlights how biological systems frequently organize information in ways that may inspire new technological approaches.
The idea of DNA-inspired computing remains largely speculative. However, the remarkable efficiency and robustness of biological information systems suggests that studying nature’s solutions may still reveal new possibilities for future computational architectures.
.:.